Reputation: 49
Adding orthogonal regression line in ggplot
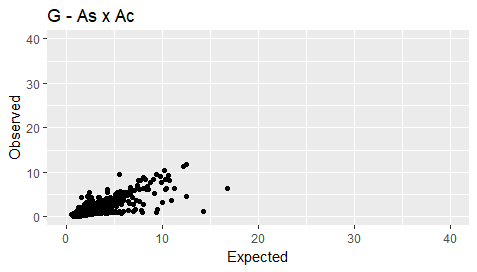
I have plotted a scatter graph in R, comparing expected to observed values,using the following script:
library(ggplot2)
library(dplyr)
r<-read_csv("Uni/MSci/Project/DATA/new data sheets/comparisons/for comarison
graphs/R Regression/GAcAs.csv")
x<-r[1]
y<-r[2]
ggplot()+geom_point(aes(x=x,y=y))+
scale_size_area() +
xlab("Expected") +
ylab("Observed") +
ggtitle("G - As x Ac")+ xlim(0, 40)+ylim(0, 40)
My plot is as follows:
I then want to add an orthogonal regression line (as there could be errors in both the expected and observed values). I have calculated the beta value using the following:
v <- prcomp(cbind(x,y))$rotation
beta <- v[2,1]/v[1,1]
Is there a way to add an orthogonal regression line to my plot?
Upvotes: 2
Views: 1367
Answers (3)

Reputation: 28371
Borrowed from this blog post & this answer. Basically, you will need Deming function from MethComp or prcomp from stats packages together with a custom function perp.segment.coord. Below is an example taken from above mentioned blog post.
library(ggplot2)
library(MethComp)
data(airquality)
airquality <- na.exclude(airquality)
# Orthogonal, total least squares or Deming regression
deming <- Deming(y=airquality$Wind, x=airquality$Temp)[1:2]
deming
#> Intercept Slope
#> 24.8083259 -0.1906826
# Check with prcomp {stats}
r <- prcomp( ~ airquality$Temp + airquality$Wind )
slope <- r$rotation[2,1] / r$rotation[1,1]
slope
#> [1] -0.1906826
intercept <- r$center[2] - slope*r$center[1]
intercept
#> airquality$Wind
#> 24.80833
# https://stackoverflow.com/a/30399576/786542
perp.segment.coord <- function(x0, y0, ortho){
# finds endpoint for a perpendicular segment from the point (x0,y0) to the line
# defined by ortho as y = a + b*x
a <- ortho[1] # intercept
b <- ortho[2] # slope
x1 <- (x0 + b*y0 - a*b)/(1 + b^2)
y1 <- a + b*x1
list(x0=x0, y0=y0, x1=x1, y1=y1)
}
perp.segment <- perp.segment.coord(airquality$Temp, airquality$Wind, deming)
perp.segment <- as.data.frame(perp.segment)
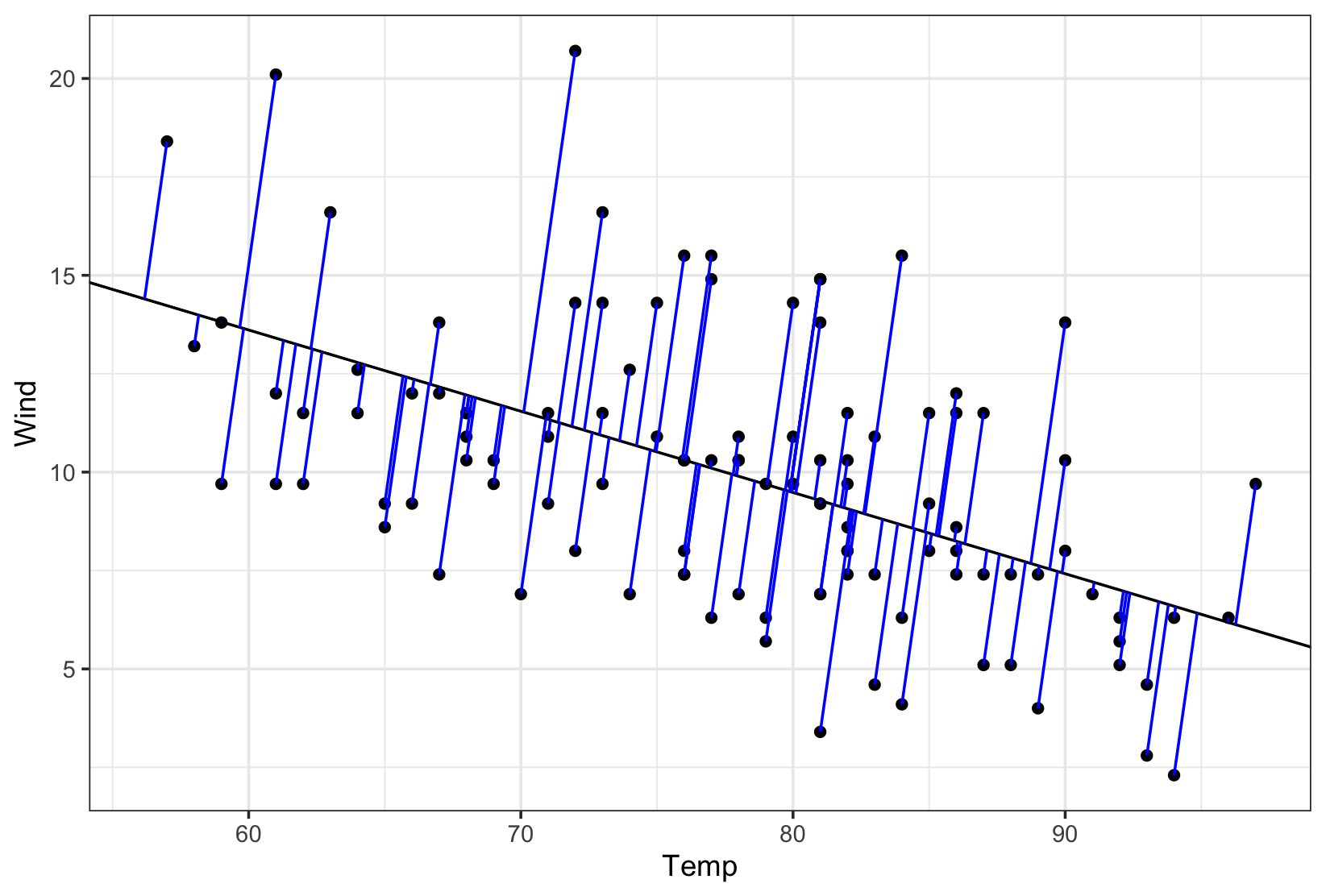
# plot
plot.y <- ggplot(data = airquality, aes(x = Temp, y = Wind)) +
geom_point() +
geom_abline(intercept = deming[1],
slope = deming[2]) +
geom_segment(data = perp.segment,
aes(x = x0, y = y0, xend = x1, yend = y1),
colour = "blue") +
theme_bw()
Created on 2018-03-19 by the reprex package (v0.2.0).
Upvotes: 3
Reputation: 9496
The MethComp package seems to be no longer maintained (was removed from CRAN).
Russel88/COEF allows to use stat_/geom_summary with method="tls" to add an orthogonal regression line.
Based on this and wikipedia:Deming_regression I created the following functions, which allow to use noise ratios other than 1:
deming.fit <- function(x, y, noise_ratio = sd(y)/sd(x)) {
if(missing(noise_ratio) || is.null(noise_ratio)) noise_ratio <- eval(formals(sys.function(0))$noise_ratio) # this is just a complicated way to write `sd(y)/sd(x)`
delta <- noise_ratio^2
x_name <- deparse(substitute(x))
s_yy <- var(y)
s_xx <- var(x)
s_xy <- cov(x, y)
beta1 <- (s_yy - delta*s_xx + sqrt((s_yy - delta*s_xx)^2 + 4*delta*s_xy^2)) / (2*s_xy)
beta0 <- mean(y) - beta1 * mean(x)
res <- c(beta0 = beta0, beta1 = beta1)
names(res) <- c("(Intercept)", x_name)
class(res) <- "Deming"
res
}
deming <- function(formula, data, R = 100, noise_ratio = NULL, ...){
ret <- boot::boot(
data = model.frame(formula, data),
statistic = function(data, ind) {
data <- data[ind, ]
args <- rlang::parse_exprs(colnames(data))
names(args) <- c("y", "x")
rlang::eval_tidy(rlang::expr(deming.fit(!!!args, noise_ratio = noise_ratio)), data, env = rlang::current_env())
},
R=R
)
class(ret) <- c("Deming", class(ret))
ret
}
predictdf.Deming <- function(model, xseq, se, level) {
pred <- as.vector(tcrossprod(model$t0, cbind(1, xseq)))
if(se) {
preds <- tcrossprod(model$t, cbind(1, xseq))
data.frame(
x = xseq,
y = pred,
ymin = apply(preds, 2, function(x) quantile(x, probs = (1-level)/2)),
ymax = apply(preds, 2, function(x) quantile(x, probs = 1-((1-level)/2)))
)
} else {
return(data.frame(x = xseq, y = pred))
}
}
# unrelated hlper function to create a nicer plot:
fix_plot_limits <- function(p) p + coord_cartesian(xlim=ggplot_build(p)$layout$panel_params[[1]]$x.range, ylim=ggplot_build(p)$layout$panel_params[[1]]$y.range)
Demonstration:
library(ggplot2)
#devtools::install_github("Russel88/COEF")
library(COEF)
fix_plot_limits(
ggplot(data.frame(x = (1:5) + rnorm(100), y = (1:5) + rnorm(100)*2), mapping = aes(x=x, y=y)) +
geom_point()
) +
geom_smooth(method=deming, aes(color="deming"), method.args = list(noise_ratio=2)) +
geom_smooth(method=lm, aes(color="lm")) +
geom_smooth(method = COEF::tls, aes(color="tls"))

Created on 2019-12-04 by the reprex package (v0.3.0)
Upvotes: 2

Reputation: 1826
I'm not sure I completely understand the question, but if you want line segments to show errors along both x and y axis, you can do this using geom_segment.
Something like this:
library(ggplot2)
df <- data.frame(x = rnorm(10), y = rnorm(10), w = rnorm(10, sd=.1))
ggplot(df, aes(x = x, y = y, xend = x, yend = y)) +
geom_point() +
geom_segment(aes(x = x - w, xend = x + w)) +
geom_segment(aes(y = y - w, yend = y + w))
Upvotes: 0
Related Questions
- Adding a regression line on a ggplot
- Add OLS squares to ggplot2 graph
- Adding Regression Line Equation and R2 on SEPARATE LINES graph
- ggplot2: How to plot an orthogonal regression line?
- Adding regression line in ggplot2
- Add quadratic regression line to existing ggplot R
- Adding a separate line of regression to ggplot in R
- Adding R squared value to orthogonal regression line in R
- ggplot2 adding regression line(logic)
- Regression line in ggplot2